In this lesson, you will learn about unsupervised machine learning algorithms—including principal components analysis, K-means clustering, a...
In this lesson, you will learn about unsupervised machine learning algorithms—including principal components analysis, K-means clustering, and hierarchical clustering—and determine the problems for which they are best suited
Unsupervised learning is machine learning that does not use labeled data (i.e., no target variable); thus, the algorithms are tasked with finding patterns within the data themselves. The two main types of unsupervised ML algorithms (displayed below) are dimension reduction, using principal components analysis, and clustering, which includes k-means and hierarchical clustering. These will now be described in turn.
Principal Components Analysis
Dimension reduction is an important type of unsupervised learning that is used widely in practice. When many features are in a dataset, representing the data visually or fitting models to the data may become extremely complex and “noisy” in the sense of reflecting random influences specific to a dataset. In such cases, dimension reduction may be necessary. Dimension reduction aims to represent a dataset with many typically correlated features by a smaller set of features that still does well in describing the data.
A long-established statistical method for dimension reduction is principal components analysis (PCA). PCA is used to summarize or transform highly correlated features of data into a few main, uncorrelated composite variables. A composite variable is a variable that combines two or more variables that are statistically strongly related to each other. Informally, PCA involves transforming the covariance matrix of the features and involves two key concepts: eigenvectors and eigenvalues. In the context of PCA, eigenvectorsdefine new, mutually uncorrelated composite variables that are linear combinations of the original features. As a vector, an eigenvector also represents a direction. Associated with each eigenvector is an eigenvalue. An eigenvalue gives the proportion of total variance in the initial data that is explained by each eigenvector. The PCA algorithm orders the eigenvectors from highest to lowest according to their eigenvalues—that is, in terms of their usefulness in explaining the total variance in the initial data (this will be shown shortly using a scree plot). PCA selects as the first principal component the eigenvector that explains the largest proportion of variation in the dataset (the eigenvector with the largest eigenvalue). The second principal component explains the next-largest proportion of variation remaining after the first principal component; this process continues for the third, fourth, and subsequent principal components. Because the principal components are linear combinations of the initial feature set, only a few principal components are typically required to explain most of the total variance in the initial feature covariance matrix.
Video here.
First and Second Principal Components of a Hypothetical Three-Dimensional Dataset
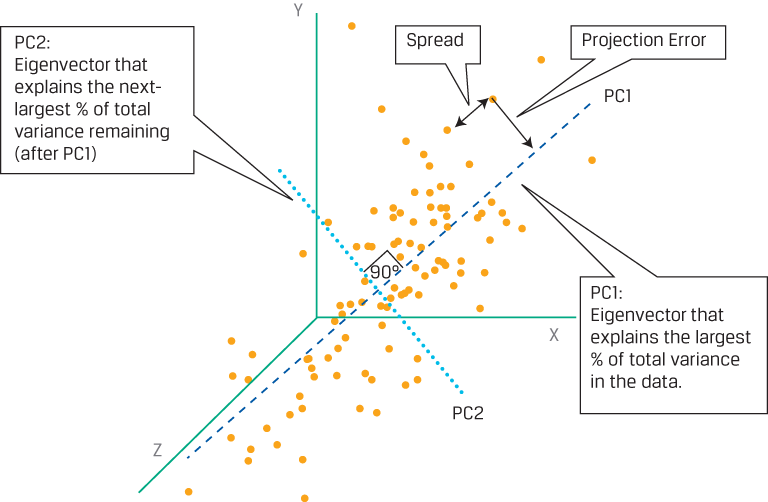
This is a hypothetical dataset with three features, so it is plotted in three dimensions along the x-, y-, and z-axes. Each data point has a measurement (x, y, z), and the data should be standardized so that the mean of each series (x’s, y’s, and z’s) is 0 and the standard deviation is 1. Assume PCA has been applied, revealing the first two principal components, PC1 and PC2. With respect to PC1, a perpendicular line dropped from each data point to PC1 shows the vertical distance between the data point and PC1, representing projection error. Moreover, the distance between each data point in the direction that is parallel to PC1 represents the spread or variation of the data along PC1. The PCA algorithm operates in such a way that it finds PC1 by selecting the line for which the sum of the projection errors for all data points is minimized and for which the sum of the spread between all the data is maximized. As a consequence of these selection criteria, PC1 is the unique vector that accounts for the largest proportion of the variance in the initial data. The next-largest portion of the remaining variance is best explained by PC2, which is at right angles to PC1 and thus is uncorrelated with PC1. The data points can now be represented by the first two principal components. This example demonstrates the effectiveness of the PCA algorithm in summarizing the variability of the data and the resulting dimension reduction.












BÌNH LUẬN